Scalable Neuroscience: Plenty of Room at the Bottom for Accelerated Returns
Google Inc.
tld@google.com
November 26, 2012
AbstractThis is the transcript of an invited lecture delivered at Stanford University on November 26, 2012. It was prompted by a growing awareness that some of the speculation regarding nanoscale machines performing useful services in fully functioning human brains might be realized sooner than originally thought possible. In this lecture I focus primarily on how synthetic biology and recombinant DNA technology might be applied to reading off the state of a living brain as an alternative to surgically implanted electrodes that only sample a tiny fraction of cells and imaging technologies such as fMRI that provide relatively poor spatial and temporal resolution and are only capable of recording a rough correlate of the signals that we really care about. Clearly interventions involving recombinant DNA modification of healthy cells in human subjects are not likely to be approved without years of technical refinement and careful testing. That said, it may make sense to redirect additional effort into exploring the options given their significant promise and the steep gradient of recent progress in the related basic science and technology. I thank Professor Shoham and his class at Stanford for encouraging me to prepare this lecture and share my observations.
 |
In 1959 Richard Feynman, gave a talk at Caltech entitled “There’s Plenty of Room at the Bottom” in which he imagined technologies that would enable us to peer at individual atoms and — perhaps more compelling to tool-using Homo Sapiens — actually manipulate atoms. He anticipated the atomic force microscope and the enormous potential inherent in the ability to explore the world at the atomic scale1. He also anticipated the development of nanotechnology capable of interacting directly with atoms and molecules and building nanoscale machines. He was particularly enamored of these “tiny machines” as he called them in a subsequent lecture and he issued several challenges for the creation of particular machines offering prize money as an incentive to aspiring inventors.
 |
Along with John von Neumann, his former colleague at Los Alamos, Feynman fully appreciated that nature had already solved the problem of atomic-scale machines, and he considered biological machines a proof that such technology was possible and only a matter of time before engineers would match or surpass natural selection. He provided insights into the challenges faced by natural and man-made tiny machines, describing how, as you descend to smaller and smaller scales, different physical laws dominate. E. Coli use corkscrew-shaped flagella and molecule-sized motors to propel themselves through a watery fluid which is for them a viscous medium as thick as molasses. For organisms operating at millimeter scales, surface tension is an important consideration; at nanometer scales, Van der Waals force starts to play a key role2.
 |
Today, we frequently see articles in the main-stream press describing advances in microelectromechanical systems or MEMS which are devices typically manufactured using semiconductor fabrication technologies and consisting of components from 1 to 100 micrometres is size. At this scale electrostatic surface effects dominate over volume effects such as inertia or thermal mass. We are also seeing new materials that are hybrids combining, for example, biologically-based substrates constructed by folding strands of DNA into three-dimensional shapes, and then adding atoms of gold or other exotic materials as conductors to implement specialized sensors and communication devices.
 |
In 1965, Gordon Moore made the observation that over the history of modern computing hardware, beginning with the invention of the integrated circuit in 1958, the number of transistors on an integrated circuit doubled approximately every two years. This exponential trend has continued more or less unabated to this day and promises to continue for some indeterminate time into the future. In this graph, the fabrication process — a modern version of lithography — referred to here as “technology nodes” — is now around 22 nanometers which allows printed lines etched on a silicon die — referred to here as “gate lengths” — of around 30 nanometers. The molecular machines in your cells — called “ribosomes” — responsible for manufacturing proteins are about 20 nanometers end to end and ribosomes are considerably more complicated machines than a single logic gate.
 |
This next graph is a little too busy for my taste but if you take the time to parse it you’ll learn some interesting facts. For instance, the size of a transistor in an Intel 8008, a microprocessor introduced in 1972, is depicted by the large bluish-purple circle and is about twice the size of a red-blood cell, and the size of a transistor in a present-day Xeon server is about half the size of an HIV virus.
 |
The challenge of scalable neuroscience is to build instruments that enable us to record the behavior of ensembles of billions of neurons at millisecond temporal resolutions where each neuron is a machine of incredible complexity, and infer from this virtual deluge of data — “tsunami” is perhaps a more apt metaphor, the function of individual neurons and predict the collective behavior of an entire brain in both its normal and pathological operating regimes.
Much of modern experimental neuroscience is based on single-cell recordings of individual neurons or multiple neurons within a small, roughly planar area of brain tissue using an array of probes arranged in a regular grid — 10 × 10 is common — that is inserted into the brain an awake animal. Ed Boyden and his team at MIT are pushing the state of the art to enable each probe in such an array to record at multiple sites along its length thereby allowing us to collect information from many neurons in a 3-D volume.
 |
Ed has pioneered methods for using robots to insert probes in experimental animals thus eliminating one source of human error and allowing precise placement under program control [17]. He is also applying optogenetic techniques — which we will discuss in a moment — that allow us to use light to both activate and silence individual neurons.
 |
As long as we have had microscopes powerful enough to resolve individual neurons, scientists have been refining methods for imaging neural tissues using specialized preparations that make neuron cell bodies stand out and utilizing ever more powerful devices, with scanning electron microscopes currently now common in academic labs. Once the tissue is prepared and an image taken, it is generally the task of a trained neurophysiologist to interpret the image and determine where one cell leaves off and another one begins. Having skilled humans in the loop, whether working with the tissue samples or interpreting images doesn’t scale, and so research labs led by Winfried Denk at Max Planck and Sebastian Seung at MIT are developing robotic devices for handling the tissue and interpreting the results of imaging [3, 15].
Unfortunately, automating the segmentation of cell bodies is more difficult than you might imagine [26, 21]. You can plainly see the leopard cub in the top sequence of images of this slide, differentiating its torso from the tree to which it clings. Segmenting the dendrites and axons in the middle row of frames is much more difficult. You may imagine individual neurons gracefully spread out in the neural tissue like free-floating seaweed fronds, but it is more accurate to imagine the neurons as spaghetti nooodles densely packed into a can. The task is made somewhat more tractable by highlighting selected neurons using color-coded fluorescent markers [14] — see Brainbow — as shown in the bottom panel, but this technology is not likely to scale due to the combinatorics involved in differentiating so many closely packed cell bodies.
 |
Sebastian’s goal is to compute the connectome — the graph of neurons and their active connections — for interesting tissue samples, starting with the retina, then a mouse brain and ultimately a human brain [25]. Even if we can improve our image processing algorithms to accurately segment cell bodies, we would still need a warehouse full of robotic tissue handlers and electron microscopes to process even a single mouse brain in a reasonable amount of time. A single cubic millimeter of neural tissue produces a petabyte of image data when scanned. One would hope there’s a better way.
 |
The method of preparing a tissue sample, slicing it into thin sections, and scanning each slice with an electron microscope that is being used to reconstruct the connectome can also be applied to determine where in the cell different proteins are utilized. Stephen Smith and his colleagues at Stanford have developed a new imaging technique they call array tomography that combines electron microscopy with immunofluorescence to visualize the distribution of specific proteins in the cell [20]. Immunofluorescence takes advantage of the specificity of antibodies to their corresponding antigens to tag proteins with fluorescent dyes so they can be imaged with a scanning electron microscope. Smith and his team have used this technique to investigate the diversity of different synapse types as identified by their characteristic protein signatures [22] and the Allen Institute for Brain Science has used similar techniques in generating data for their incredibly useful Brain Atlas resources.
 |
To get a better idea of the scale of the problems we’re considering, here are some numbers that quantitative neuroscientists keep in mind when doing back-of-the-envelope calculations. 100 billion of anything is a lot, but 100 billion sophisticated computing machines is staggering. White matter consists mostly of glial cells and myelinated3 axons that covered with an insulating sheath that speeds transmission and ameliorates the effects of noise and the potential for crosstalk.
 |
The number of neurons is perhaps less important than the number of active connections or synapses. Scott McNealy at SUN Microsystems was fond of saying “It’s the network stupid”, and his statement applies to computing in the brain as well as computing networks that characterize modern cloud computing architectures. There are something on the order of 1000 trillion synapses in a human brain and the molecular machinery operating at these connections is similarly complex.
 |
Certainly we are not going to position a probe at every location within a few microns of a synapse in the human brain. We may, however, be able to develop nanoscale machines and distribute them throughout the brain so that every neuron can comfortably accommodate a few thousand of these machines positioned strategically in its active synapses. These machines would be designed to record the passage of proteins called neurotransmitters, which are the primary currency for exchanging information between neurons. Fortunately, given the state of the art in molecular biology, we don’t have to generate these machines de novo, but rather it seems plausible that we will be able to adapt existing cellular machinery from a variety of organisms to perform the basic sensing and communicating tasks required.
Every cell in the human body, and neurons in particular, contains a collection of molecular machines, some free floating and independent, and others anchored in cellular structures called organelles that serve as factories for the production and shaping of proteins, lipids and other macromolecules. In this graphic, the structure labeled (2) is the nucleus of the cell containing the DNA instructions to build the entire organism. (1) is called the nucleolus and is responsible for transcribing (ribosomal) RNA used to build ribosomes — shown as small dots one of which is labeled (3) — that are housed in the endoplasmic reticulum (5) a structure that serves to fold the newly minted but only partially-formed proteins into their final conformations. There are also organelles, the Golgi apparatus (6), that manufacture the many types of membranes that make up the structural members of the cell and a class of organelles called mitochondria (9) which are actually symbiotic organisms with their own separate DNA that set up housekeeping within the cells of eukaryotes many millions of years ago and made themselves indispensable.
 |
All of these molecular machines are constantly at work performing the specialized functions of the particular cell type as well as performing a range housekeeping and routine repair chores. Everything that happens in the cell depends on the manufacture of proteins and amino acids using the cell’s DNA — its genetic code — as a blueprint. Indeed, the genetic code is being read and decoded at every moment in every cell in your body. You can’t move a muscle or think a thought without the production of scores of proteins and amino acids that work together to produce a dizzying array of behaviors manifest across an impressive range of temporal and spatial scales. Here’s an artistic rendering of a ribosome showing its two major molecular structures and a hint at its complicated shape and function.
 |
Ribosomes are molecular machines constructed from ribosomal RNA and additional proteins that work in concert with transfer RNAs to pluck amino acids out of the fluid or cytoplasm that comprises much of the cell’s interior and consists of water and various dissolved molecules. These amino acids are formed into polypeptide chains that are subsequently folded to produce the specific three-dimensional shape or conformation that determines the protein’s function.
 |
There are also molecular machines that serve to transport proteins from their place of manufacture, primarily organelles in a region called the soma in a neuron, to distant locations in the axons and dendrites that are involved in transferring information between neurons. Molecules called kinesins transport these protein products along pathways called microtubules.
 |
They perform this essential service to the cell by utilizing molecules of stored energy to flex the kinesins, thus altering their conformation and enabling kinesins to essentially walk along the microtubules carrying their protein cargoes.
 |
It is said that if you can imagine an operation that might be performed in a cell involving some manipulation of proteins or amino acids, then there is almost certainly some organism whose cells routinely perform that operation. Natural selection has had billions of years in which to explore the possibilities and seldom misses a trick. Molecular biologists are amassing a great catalog of such machines, many of which can be adapted to perform functions in cells other than those in which they are found naturally. This means that often as not if we need a molecular machine to perform a particular function we can order one from this catalog and adapt it to suit our purposes.
A good example of such adaptation comes from the latest generation of devices for sequencing DNA. The Human Genome Project more or less completed the sequencing of the human genome in 2004 after more than a decade of work and a cost of nearly 3 billion dollars. But this was just a single instance of the genome — actually it was a patchwork of pieces of DNA from several individuals, and, while we share a good deal of our individual genetic code, it is the differences between individuals that are likely to provide the clues in finding the causes and cures for many diseases. The race was on to drive down the cost and reduce the time required to days if not minutes.
The early gene sequencing technology required a good deal of machinery to automate what scientists had done on a smaller scale at their lab benches using diverse reagents and complex preparations. Some of the companies offering genome sequencing imagined building warehouses full of computers and robots programmed to carry out biochemical assays. It is worth noting that every cell in your body is sequencing your genome every minute of every day. How hard could it be? Several companies in an effort to scale sequencing are working to reduce the most time consuming part — reading off the sequence of nucleotides4 in a single strand of DNA, to a process that can be carried out on a silicon chip.
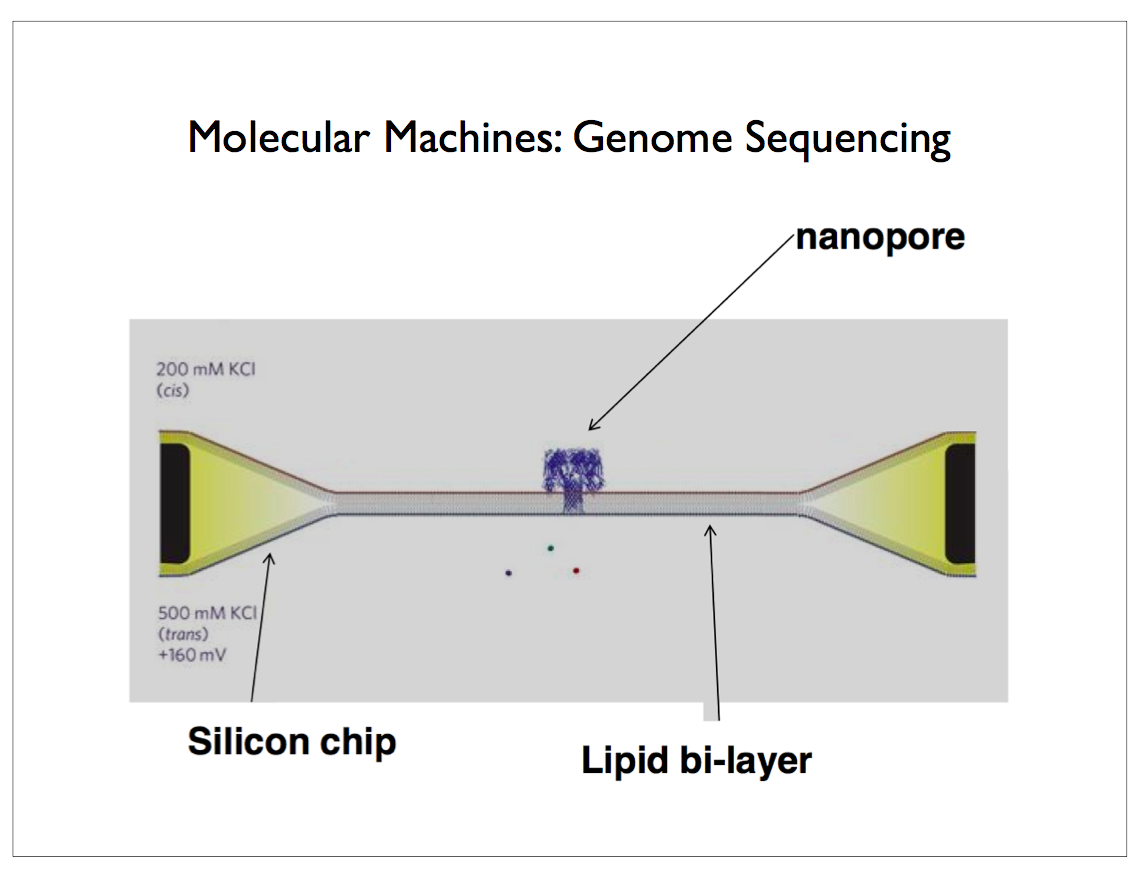 |
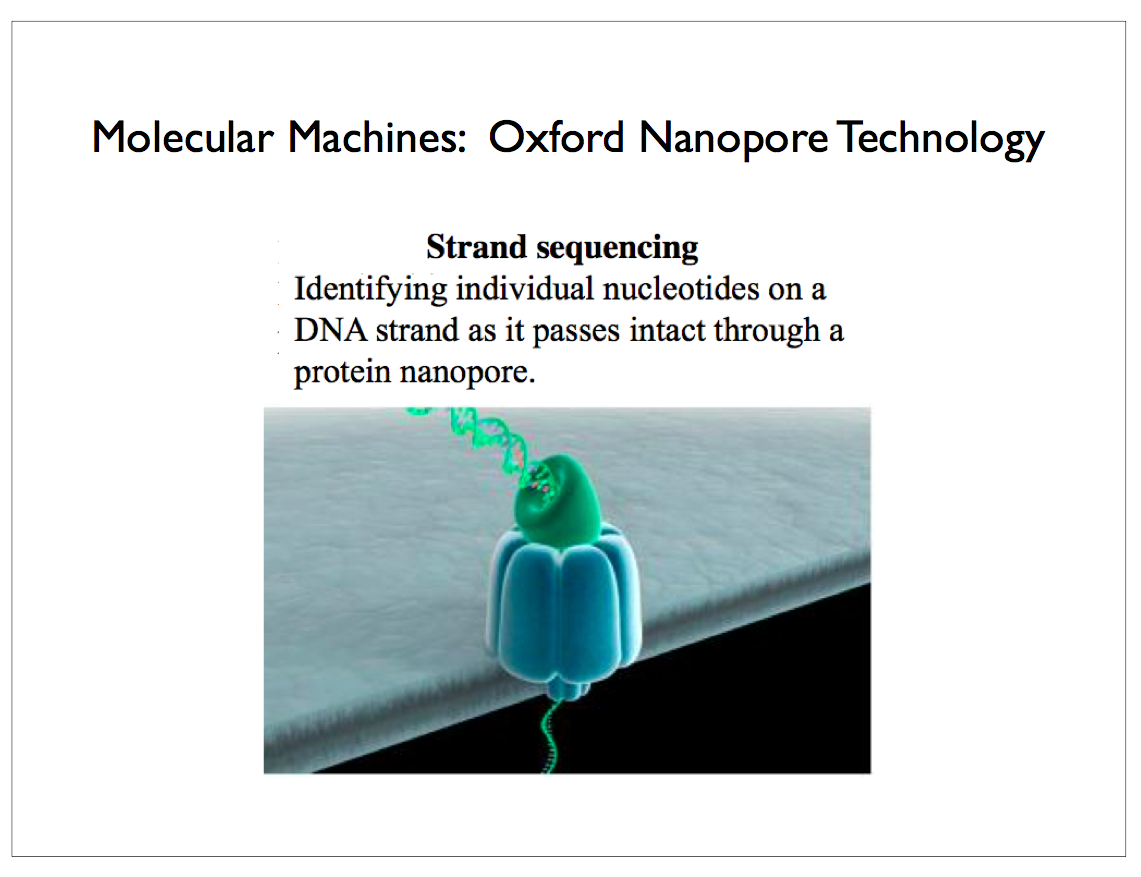
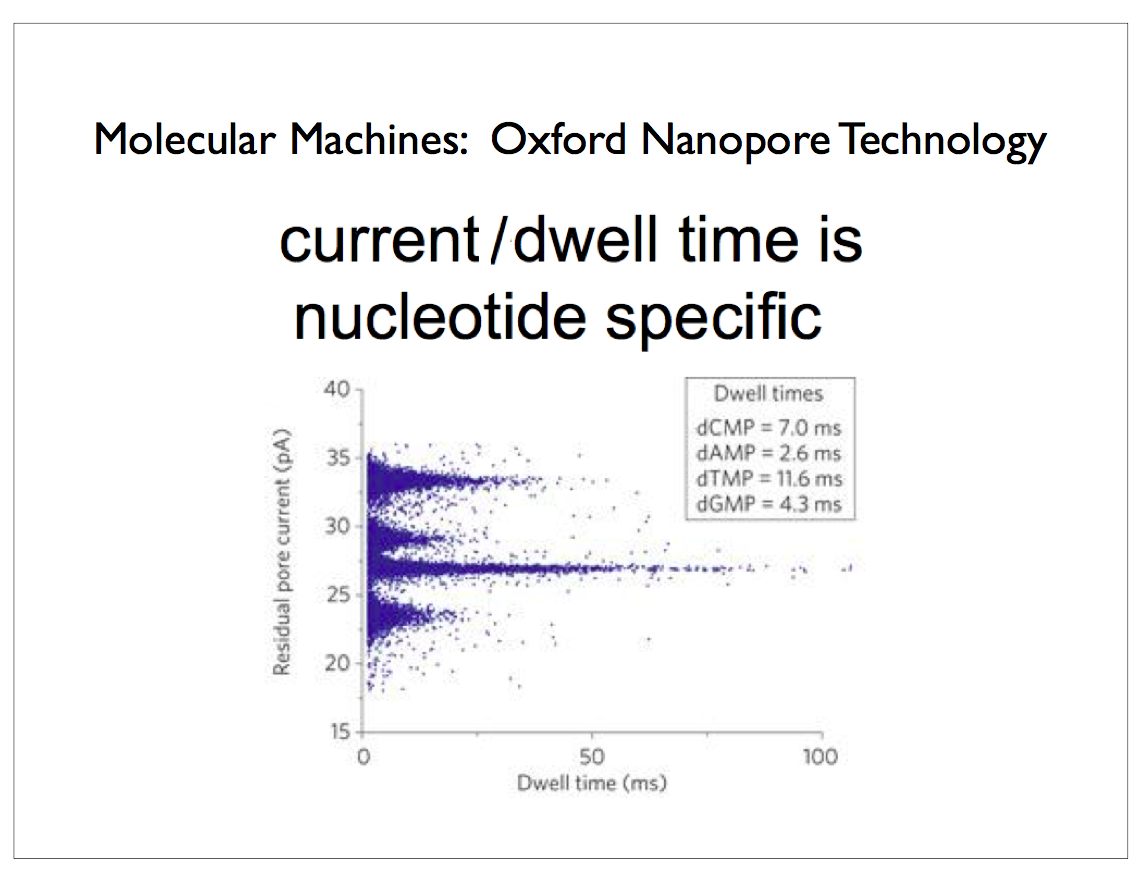
Oxford Nanopore has developed a technology that inserts a protein channel or pore in a silicon substrate. Single-strand DNA is threaded through this pore and a sensor reads out the voltage resulting from molecular changes as each nucleotide passes through the sensor. The voltage is correlated with the type of nucleotide — one of adenosine, cytidine, guanosine or thymidine, abbreviated as A, C, G and T in the case of DNA, so that the sequence can easily be read off. This scales so that you can populate a single silicon die with thousands of these pores arranged in a regular grid. It is also possible to sculpt a tiny hole in the silicon with a laser to achieve the essential properties of the protein required for sequencing. Aside from preparing and cloning the original sample of DNA which can be accomplished using PCR, this technology avoids most of the complicated chemical processes and expensive reagents that plagued earlier technologies5.
 |
 |
Another example of re-purposing existing molecular machinery comes to us from recombinant DNA technology associated most closely with DNA cloning. In one of the simpler methods applications of recombinant DNA technology, a target piece of DNA is assembled in a self-replicating package called a plasmid and introduced into an easy-to-grow bacterial host such as E. Coli. The host cells containing the plasmid are then grown in a nutrient medium using the host’s reproductive machinery to produce a large quantity of the modified organism.
 |
Francis Crick — of Crick and Watson, double-helix fame — challenged neuroscientists to determine the connectivity pattern of a single neuron and all of its neighbors. This is a special case of a connectome which we mentioned earlier. To accomplish this feat, researchers applied another recombinant DNA tool to infect progenitor stem cells that can grow into neurons with a self-replicating virus called a retrovirus. A retrovirus is an RNA-based virus that replicates in a host cell using one of its own enzymes called reverse transcriptase to produce DNA from its RNA genome. The viral DNA is then incorporated directly into the host’s genome using another enzyme called integrase. The virus thereafter replicates as part of the host cell’s DNA.
 |
HIV and rabies are particularly nasty instances of retroviruses that have been adapted for (synthetic) recombinant DNA purposes. These viruses are loaded with a DNA payload which they splice into the host’s DNA and thereby use the host’s sequencing and protein-transcribing machinery to reproduce the virus or carry out any other process that can be accomplished by the cellular machinery. In the case of addressing Crick’s challenge, the virus uses the microtubule-based transport system we described earlier to propagate copies of itself throughout the cell and infect adjacent cells connected through axonal processes. Fluorescent markers attached to the viral DNA are used to highlight the connectome and produce the images you see here.
 |
Retroviruses and recombinant DNA technology also provide powerful tools for controlling individual neurons. Optogenetics is the name given to a collection of tools that use light to activate or silence neurons [4, 5]. The electrical properties of neurons and the action potentials that propagate electrical signals along axonal processes are governed by differences in ion concentrations across the cell membrane. Voltage- and ligand-gated ion channels — a ligand is a biomolecule involved in cellular signalling and neurotransmitters constitute the class we are most concerned with here — serve to alter ion concentrations by allowing specific ions — sodium, potassium and calcium — to flow through the membrane and reestablish the equilibrium in accord with the prevailing voltage and concentration gradients. Our cellular machinery is exquisitely tuned so that only a few ions are required to cause substantial changes in the voltage difference across the axonal membrane.
 |
Some species of algae have light-gated ion channels that can be introduced into neurons using recombinant DNA technology. If you shine a light of the proper frequency on these channels they open, altering the potential across the cell membrane and, depending on the voltage drop, initiating an action potential to propagate along the the axon. There is another technique in the optogenetics toolkit that relies on a different wavelength of light to silence a neuron, thus preventing it from initiating a spike, and thereby enabling the researcher to both turn on and turn off individual neurons. This approach to controlling neurons is extremely useful in establishing cause-and-effect relationships, but it still requires additional instrumentation, such as conventional voltage-sensing probes to measure electrical activity in neurons.
 |
The use of tame variants of naturally occurring retroviruses is a powerful tool for co-opting cells to perform tasks required for the production and delivery of drugs and gene therapies. The field of synthetic biology focuses on engineering biological circuits6 consisting of standard components that can be ordered from a catalog and depended upon to reliably produce a specified behavior. Ultimately, the goal is to build molecular machines using such components so that all the operations required for a given application are performed entirely within our living cells [10].
 |
In the case of the nanopore gene sequencing technology, a sample is extracted from the target organism and the sequencing is performed externally. In the case of tracing the connectome using a rabies virus, the experiments are performed in vitro using cells grown in a culture. How might we operate directly on cells in vivo, that is to say in live animals without requiring surgery, inserting probes or sacrificing the animal to extract information for analysis.
One possible answer is to use recombinant DNA technology to re-program cells to perform new operations while not interfering with their normal function. Where in the case of employing a retrovirus to trace the connectome we used the existing inter-cellular transport mechanism, in the case of delivering a viral vector to a large population of target cells — neurons in our case — we might exploit the system of arteries and capillaries that deliver nutrients and oxygen to every cell in the human body. In the particular case of the brain, delivery is complicated by the intervention of the blood-brain barrier which consists of a membrane called the endothelium that surrounds each capillary and through which every molecule entering the brain must pass. Once inside the brain glial cells called astrocytes assist in the exchange of oxygen and glucose and the production of enzymes and neurotransmitters essential for neurons to perform their duties.
 |
The blood brain barrier protects the brain from toxins and pathogens that might disrupt the neural machinery controlling vital processes throughout the body. Unfortunately for those affected by viral-borne brain diseases, nature has figured out how to bypass the barrier and the HIV and rabies retroviruses mentioned earlier are examples of such pathogens. The silver lining is that we are figuring out how to use these same viral vectors to repair cell damage and deliver drug payloads selectively to targets throughout the body and the brain in particular. We’re also figuring out ways to foil natural viruses so they can’t cross the blood-brain barrier.
 |
If we can use the arteries and capillaries to distribute and deliver molecular machines, then we might also use the lymph system and the vessels that return blood to the lungs and heart and waste products to the kidneys as a means of conveying information to locations external to the central nervous system where it might be more easily processed, say, using an artificial filtering process akin to dialysis7. This would provide an expedient for reading out neural-state information in lieu of more complicated nanotechnology solutions employing tiny radio transmitters that have been suggested in the literature.
Now we have a means of providing input and extracting output from the brain9. Granted that the input modality we’ve been exploring requires we infect each cell with virus and modify its DNA, and that would likely not serve as the input side of a real-time computer interface. On the output side however, we might have a shot at being able to observe a behaving brain at an unprecedented scale and level of detail. Assuming that our virally-delivered molecular machines don’t interfere with the normal operation of the cells, we could in principle develop technology for reading off states of the brain that would not harm the host and could operate indefinitely10. What sort of information might we want to collect and how would we go about doing so?
Traditionally the focus has been on recording spike trains in the form of changes in the membrane potential of individual neurons. However, the signaling pathways in the brain are subtle and multitude; they include electrical pathways12 in the form of action potentials and voltage-gated ion channels, genetic pathways in the form of DNA translated into RNA and proteins expressed and transported within the cell, and chemical pathways in form of neurotransmitters which are emitted into the synaptic cleft separating an axon and a dendrite and serve to open ligand-gated ion channels on the dendrite.
Here we see a schematic synapse showing the neurotransmitters packaged in vesicles (2) in the pre-synaptic neuron A, ligand-gated ion channels (5) in the post-synaptic neuron B, and a mitochondrial organelle (1) that provides energy in the form of ATP. Other components include voltage-gated calcium13 channels (6) that are activated by action potentials and cause the vesicles to merge with the cell membrane and neurotranmitters to flood into the synaptic cleft and additional machinery responsible for scavenging neurotransmitters in a process called reuptake.
 |
Must we record the state of all these components in order to obtain a complete picture? Perhaps, but it may be that the proteomic history — the record of specific proteins expressed and transported across synapses — is sufficient to infer most of what is going on informationally and computationally within the brain. In any case, we are going to assume so for the remainder of this discussion, and make the additional simplifying assumption that the production and transfer of neurotransmitters provide enough information.
Our grand goal is to collect enough data to infer not only the structure and circuitry of the brain — what we have been calling the connectome, but the function of smaller, anatomically-localized neural circuits, larger super-complexes of neurons that implement functional areas such as the visual cortex, and the recurrent pathways linking these functional areas to support high-level cognition. We hope to abstract the behavior of these diverse neural circuits and build models to test our understanding. Our progress so far suggests that we will have to record simultaneously from large collections of neurons to make additional progress on these challenging problems.
 |
Here’s a very rough sketch for how we might record the proteomic history of a behaving brain. First off we need to be able to identify what neurotransmitter is being conveyed, which neuron is transmitting the information and which neuron is receiving it. Essentially we need a unique identifier for each class of neurotransmitter and each individual neuron. A recent paper in Nature described a scalable method for generating self-assembling barcodes using DNA origami as a substrate and fluorescent tags to encode the digital information [19]. Something like this method might suffice to encode the unique identifiers that we require.
 |
Next we need to associate neurotransmitters with their identifying barcodes and convey these barcodes along with the neurotransmitters making sure that they find their way into the receiving neuron where they can be assembled into packets that describe each event as a triple of three barcodes encoding the transmitting neuron, the receiving neuron and the class of neurotransmitter conveyed. Once assembled these packets would be flushed into the cerebrospinal fluid to be subsequently eliminated from the brain via the lymph and blood circulation system.
Fleshing this out would require a great deal of speculation, and so to reduce the hand waving to a tolerable level let’s consider one more-or-less concrete proposal suggested in a paper [28] out of Tony Zador’s group at Cold Spring Harbor Laboratory. The authors of this paper propose the idea of sequencing the connectome in their paper of the same title. They break down the problem into three components: (a) label each neuron with a unique DNA sequence or barcode, (b) propagate the barcodes from each source neuron to each synaptically-adjacent sink neuron — this results in each neuron collecting a “bag of barcodes”, and (c) for each neuron combine its barcodes in source-sink pairs for subsequent high-throughput sequencing.
 |
It would seem the authors have in mind sacrificing the animal in the final step, but it may be possible to pass source-sink pairs through the cell membrane and into the lymph-blood system for external harvest as proposed earlier. They suggest that propagation might be accomplished using a trans-synaptic virus such as rabies [23]. Additional techniques would be required to map barcodes to brain areas. The authors claim to be developing an approach based on PhiC31 integrase for joining barcodes and PRV amplicons [24] for trans-synapatic barcode propagation. While quite ambitious with many complicated steps yet to be filled in, a number of neuroscientists, myself included, believe that some variant of this idea could be accomplished in the relatively near-term future, and there are plans afoot [2] to take on even more ambitious goals14.
I could literally go on for hours and have in some more intimate classroom situations. I’ve created an annotated transcription of this talk with lots of footnotes providing additional detail and relevant papers for your further reading. I believe wanting to tell everyone about what you’ve discovered is a wonderful child-like characteristic that is also incredibly valuable to both the individual and society. It is a form of public thinking that is under-appreciated and often discouraged in precocious children. I encourage it in everyone I meet and cherish it in my students and colleagues. I thank you for your patience indulging me in my habit.
References
1 It helps to get some appreciation of scale by comparing the sizes of objects that we can experience. For example, the height of Mount Everest is 29,029 feet (8,848 meters). We can put that in human perspective by a simple change in the units that we use for our measurements. Mount Everest is about 5,000 times as high as a six foot person — (/ 29029.0 6.0) = 4838.17. The same method of contrast works for smaller objects at the nanoscale. The cell body or soma of a neuron can vary between 4 and 100 microns. A six foot person (1.8288 meters) is roughly 100,000 times the size of a neuron cell body — (/ (* 1.8288 (expt 10.0 6.0)) 10.0) = 182880.0.
In dealing with irregularly shaped objects, we use idealizations such as asssuming that the sun and earth are spherical. The radius of the earth is around 6,371 kilometers — (defconst earth-radius 6371.0). The radius of the sun is around 696,000 kilometers — (defconst sun-radius 696000.0). The radius of the sun is around 100 times the earth’s radius — (/ 696000.0 6371.0) = 109.25. The volume of the sun is around 1,000,000 times the earth’s volume. If you’re curious about the odd parenthetical expressions, I’m writing these notes using the Emacs editor and using its scripting language — a dialect of Lisp — to perform my simple back-of-the-envelope calculations:
(defconst float-pi 3.141592653589793 "The value of Pi.") (defun volume-of-sphere (radius) (* (/ 4.0 3.0) float-pi (expt radius 3.0))) (defun ratio-of-volumes (radius-1 radius-2) (/ (volume-of-sphere radius-1) (volume-of-sphere radius-2))) (defconst sun-radius 696000.0 "Radius of the sun in kilometers.") (defconst earth-radius 6371.0 "Radius of the earth in kilometers.") (ratio-of-volumes sun-radius earth-radius) ;; => 1,303,781.78 (defconst solar-system-radius 5913520000.0 "Radius of the solar system in kilometers.") (ratio-of-volumes solar-system-radius sun-radius) ;; => 613,352,996,129.95
Measuring the size of the solar system requires that we introduce additional assumptions regarding the shape of objects whose boundaries are inconstant. The radius of the solar system is taken to be the average distance between the Sun and Pluto is 5,913,520,000 kilometers — (defconst solar-system-radius 5913520000.0). The radius of the solar system is around 10,000 times the sun’s radius — (/ 5913520000.0 696000.0) = 8496.44. The radius of the solar system is around 1,000,000 times the earth’s radius — (* 8496.44 109.25) = 928236.07.
Comparing the relative sizes of objects at scales below the nanoscale, e.g., electrons and protons, presents new challenges. A proton is not an elementary particle and hence it possesses a physical size, although its spatial envelope varies since the surface of a proton is somewhat fuzzy due to being defined by the influence of forces that don’t come to an abrupt end. The proton is about 1.6-1.7 femtometers in diameter. Note that one femtometer is 1.0 × 1015 meters or 0.001 picometer, and one nanometer is 1,000 picometers or 1,000,000 femtometers.
An electron is an elementrary particle and hence it is described in quantum mechanical terms as a wavefunction, which in principle covers all space. There is a measure called the classical electron radius, also known as the Lorentz radius which is roughly the size the electron would need to have for its mass to be completely due to its electrostatic potential energy — not taking quantum mechanics into account.
If we consider the size of atoms, our measurements are a little easier to describe but still require we deal with some degree of ambiguity at the quantum level. The bond length is the average distance between the nuclei of two bonded atoms, e.g., the bond length between a carbon and a hydrogen atom is around 100 picometers or 0.1 nanometers. The atomic radius is the mean distance from the nucleus to the boundary of the surrounding cloud of electrons, e.g., atomic radius of hydrogen is 25 picometers and that for carbon is 70 picometers.
2 When Feynman discussed assembling nanoscale machines, he would often speak of first building a set of 1/4 scale tools, using them to build 1/16 scale tools, and so on, ultimately constructing millions of entire nanoscale factories. This leads some to think that nanoscale assembly will look like macroscale assembly — tiny machine tools made out of rigid parts, constructing nanoscale products out of materials that behave like the materials we encounter in everyday life. If we were to proceed with this intuition, we would very likely end up being disappointed. Nanoscale fabrication and assembly present new engineering challenges precisely because different physical laws dominate at different scales, but it also offers powerful new opportunities for combinatorial scaling.
Objects at the nanoscale, organic molecules in the case of biological systems, tend to be “flexible”, “sticky” and perpetually “agitated.” “Flexibility” refers to the fact that proteins and other large molecules that comprise biological systems generally have multiple shapes or “conformations”. Even once proteins are folded into a particular conformation, the geometric arrangement of their constituent atoms changes in accord with the attractive or repulsive forces acting between parts of the protein, e.g., Van der Waals force, and interactions with other molecules in their vicinity, e.g., due to the forces involved in making and breaking covalent bonds.
“Stickiness” refers to the fact that these molecules routinely exchange electrons allowing new molecules to be formed from existing molecules by way of chemical reactions catalyzed by enzymes. These molecules have locations — the “sticky” sites — corresponding to molecular bonds where electrons can shift their affinity to create new bonds with other nearby molecules — and hence “stick” together. Finally, “agitated” refers to the fact that the molecules are constantly in motion due to changes in conformation, interaction with other macromolecules, and being struck by smaller fast moving atoms and molecules. The attendant forces cause individual particles to undergo a random walk, with the behavior of the ensemble as a whole referred to as brownian motion.
In nanoscale engineering, these properties of nanoscale objects can be channeled to create products by self-assembly. The study of soap films provides a relatively simple introduction to the natural processes involved in self-assembly, and there are a number of popular books in the library that detail these same processes at work in biological systems [11, 16]. Physicists like to joke that you don’t study quantum mechanics to understand it — since that is clearly impossible, only to apply it — and, of course, that implies a book on quantum theory that doesn’t include a lot of worked-out examples and derivations is of little value. Quantum mechanics is definitely a prerequisite for many nanoscale engineering applications, but it is also necessary to acquire intuitions that enable us to imagine how molecules interact both in pairs and in larger ensembles at these unfamiliar scales. Fortunately, biology provides us with a diverse collection of molecular machines we can study to develop those intuitions.
3 Myelin is a dielectric (electrically insulating) material that forms a layer, the myelin sheath, usually around only the axon of a neuron. It is essential for the proper functioning of the nervous system. It is an outgrowth of a type of glial cell (source)
4 Nucleotides are “biological molecules that form the building blocks of nucleic acids (DNA and RNA) and serve to carry packets of energy within the cell (ATP).” This useful graphic compactly illustrates the structural characteristics of the family of nucleotides, highlighting the phosphate groups that assist in providing the energy required for reactions catalyzed by enzymes:
 |
5 Here’s an excerpt from Oxford Nanopore’s promotional material describing their basic technology:
A nanopore is, essentially, a nano-scale hole. This hole may be:Richard A. L. Jones the author of Soft machines: nanotechnology and life [16] has some interesting observations concerning the gene-sequencing technology being developed by Oxford Nanopore in this article.
Biological: formed by a pore-forming protein in a membrane such as a lipid bilayer
Solid-state: formed in synthetic materials such as silicon nitride or graphene, or
Hybrid: formed by a pore-forming protein set in a synthetic material.
This diagram shows a protein nanopore set in an electrically resistant membrane bilayer. An ionic current is passed through the nanopore by setting a voltage across this membrane.
If an analyte passes through the pore or near its aperture, this event creates a characteristic disruption in current. By measuring that current, it is possible to identify the molecule in question. For example, this system can be used to distinguish between the four standard DNA bases G, A, T and C, and also modified bases. It can be used to identify target proteins, small molecules, or to gain rich molecular information, for example to distinguish the enantiomers of ibuprofen or molecular binding dynamics.
6 Such circuits can be used for a variety of purposes including biology-based computing. There is another, direct application of DNA to computing which was developed by Len Adleman and first applied to solving instances of the NP-complete Hamiltonian Path Problem: given an undirected graph G determine if there is a path through G that includes every vertex in G exactly once. Adleman’s original work [1] on so-called DNA computing developed into a new field which has come to be called Biocomputing.
In Genesis Machines: The New Science of Biocomputing, Martyn Amos provides an interesting description of Adleman’s work and, in particular, his application of PCR. Adleman’s DNA-based algorithm works by harnessing the ability of bacteria to replicate quickly in order to generate a large number of DNA sequences and then searching through these sequences in parallel to determine if there exists a sequence coding for a Hamiltonian path. The Amos account includes a short biography of the eccentric Kary Mullis who is credited with inventing PCR and the recipient of a Nobel prize in chemistry for his accomplishments.
8 Here is an excerpt from Ventola [27] discussing how nanoparticles — denoted “NP” in the following — are removed from circulation by the immune system; the abbreviation “RES” denotes the reticuloendothelial system which is the part of the immune system consisting of phagocytes located in reticular connective tissue and is referred to as the mononuclear phagocyte system in modern medical texts:
NPs are generally cleared from circulation by immune system proteins called opsonins, which activate the immune complement system and mark the NPs for destruction by macrophages and other phagocytes. Neutral NPs are opsonized to a lesser extent than charged particles, and hydrophobic particles are cleared from circulation faster than hydrophilic particles. NPs can therefore be designed to be neutral or conjugated with hydrophilic polymers (such as PEG) to prolong circulation time. The bioavailability of liposomal NPs can also be increased by functionalizing them with a PEG coating in order to avoid uptake by the RES. Liposomes functionalized in this way are called “stealth liposomes.”NPs are often covered with a PEG coating as a general means of preventing opsonization, reducing RES uptake, enhancing biocompatibility, and/or increasing circulation time. SPIO NPs can also be made water-soluble if they are coated with a hydrophilic polymer (such as PEG or dextran), or they can be made amphophilic or hydrophobic if they are coated with aliphatic surfactants or liposomes to produce magnetoliposomes. Lipid coatings can also improve the biocompatibility of other particles.
Relevant to the elimination of NPs by the kidneys, this paper by Choi et al [9] claims to have “precisely defined the requirements for renal filtration and urinary excretion of inorganic, metal-containing nanoparticles”, and, while somewhat narrowly focused, it provides some useful general information regarding renal filtration.
7 An even simpler expedient might involve creating information capsules that would directly marshal the normal filtration capabilities of the kidneys8 to flush the data-laden cargo into the urinary tract where it would be easier to process. In the case of lab animals like mice, a catheter could be used to collect the urine, or, simpler yet, use a nonabsorbent bedding material in the animal’s cage with a removable screened collection tray.
9 If I had to bet, I’d put my money on quantum mechanics playing a role in the development of practical methods for reading off neural states at scale. In the near-term, we may be able to utilize existing cellular transport machinery to extract neural state, but such a primarily-biological approach won’t offer high-enough temporal or spatial resolution for sophisticated brain-computer interfaces. Quantum mechanical principles such as quantum tunneling are critical in the design of semiconductors, including transistors consisting of single atoms, and technologies based on quantum dots offer efficient approaches for encoding and transporting information locally and are likely to figure in the development of nanoscale communications networks [6].
My high school physics class didn’t cover any quantum theory but I picked up a little in the electrical engineering courses I took in college. (I was a math major and so I didn’t take the full EE curriculum which I now regret.) If you weren’t exposed to quantum theory in high school or college, but know basic classical electromagnetic theory, you might want to at least learn a few quantum mechanical principles so you’ll have some clue when they come up in relation to technologies for neural interfaces.
I suggest trying to get an AP Physics Exam B level of understanding that covers Max Planck’s analysis of black-body radiation, Albert Einstein’s interpretation of the photoelectric effect, and Werner Heisenberg’s uncertainty principle along with Hermann Weyl’s more formal equation relating the standard deviation of position σx and the standard deviation of momentum σp shown here, σxσp = h/4π, for a special case involving Gaussian distributions where h is Planck’s constant.
I admit that trying to understand quantum theory is difficult and good intuitions are hard to come by — you might have heard that Einstein, who along with Planck, Heisenberg, and Niels Bohr helped to develop quantum mechanics, was not comfortable with the theory. I’ve had some success suggesting that students look at Michael Fayer’s Absolutely Small: How Quantum Theory Explains our Everyday World [13] for an account that is not only accessible but also reasonably detailed, or his textbook [12] for a more rigorous quantitative treatment. I always feel a little more comfortable with equations when I can translate them into code and perform my own synthetic experiments by playing with the constants. Here’s a Matlab implementation of Planck’s equation for calculating the electromagnetic radiation emitted by a black body in thermal equilibrium at a definite temperature.
11 High temperatures damage cells by destroying organic molecules such as proteins, carbohydrates, lipid and nucleic acids:
Alteration of Cell Walls and Membranes:
Cell Walls: Prokaryotic bacteria, eukaryotic plants and some fungi have cells that are surrounded by a protective cell wall composed mainly of structural carbohydrates and which helps maintain cell shape. Heat can disrupt the bonds within cell walls making them weak and structurally unsound.
Cell Membranes: Phospholipids are the main component of cell membranes and, in eukaryotic cells, phospholipids create an entire transport system for moving materials into, out of and around within the cell. Phospholipids become more fluid when heat is applied, disrupting the integrity of cellular membranes.
Viral Membranes: Some viruses, those that are considered to be enveloped, are surrounded by phospholipids that they steal from the cells that they parasitize. Enveloped viruses can be rendered harmless when their viral envelope is destroyed, because the virus no longer has the recognition sites necessary to identify and attach to host cells.
Damage to Proteins and Nucleic Acids:
Cellular Proteins: These large three dimensional molecules are composed of amino acids linked together by peptide bonds. Heat denatures — changes the shape of — proteins, and the 3-D structure of a protein is essential to its function. If a protein’s shape is irreversibly changed, the protein is no longer functional. Denaturation of protein is irreversible.
Nucleic Acids: Composed of linked nucleotides, nucleic acids, such as DNA and RNA contain the code for building of protein molecules. Like proteins, nucleic acids are very heat sensitive. High temperatures can result in fatal mutations to DNA or can halt the process of protein synthesis, by damaging RNA.
10 There are technical books on in vivo nanoscale communication networks [6] including discussions of the suitability of various network topologies and variant signal-transmission technologies from magnetic resonant coupling [8] to exploiting existing cellular transport and signaling pathways [18, 7]. For example, here’s a nanoscale radio receiver made from a single carbon nanotube, and a discussion of resonant inductive coupling as an efficient method for communicating over short distances.
Whether you encode information in cellular waste products or transmit the information over a local nanoscale communication network, thermodynamics dictates that you will expend energy. You either have to provide this energy locally, making less available to the cell or you have to transport energy into the cells from an external source. If the signal or power transmission involves electromagnetic radiation, then you have to be careful to avoid damaging the organic components and you will have to dissipate any waste heat11 since inevitably the operations will not be perfectly efficient.
Here’s a back-of-the-envelope calculation that you might be able to carry out if you know something about cellular and wireless technologies. Suppose you want to place a nanoscale transmitter — ultimately the gamers would like to transmit and receive but scientists are currently most interested in recording what’s going on — either inside or within a few nanometers of nearly every neuron in the primate cortex, that’s about 10 billion give or take an order of magnitude. Each transmitter would have to transmit something on the order of 40K bits per second to capture the information encoded in an action potential. A complete action-potential cycle takes around 4 milliseconds consisting of about 2 milliseconds for polarization and depolarization of the axon cell membrane followed by a refractory period of about 2 milliseconds during which the neuron is unable to fire.
 |
Existing biological systems use highly energy-efficient processes. Excerpting from a recent survey article by Mark Leeson [18]: “Recent measurements of reaction energies give values of ~10-19 J for a few hundred molecules for communication. In comparison, for CMOS, the switching energy is related to the capacitance and the square of the supply voltage. Employing 0.18 μm CMOS at 1.2 V, with an oxide thickness of 2 nanometers, the switching energy is ~10-15 J. It therefore seems likely that molecular communication mechanisms will be able to undertake computation functions that dissipate less power than current electrical components. [...] However, information propagation speeds in molecular communication are only in the hundreds of bits per second range because of the diffusion mechanism and the energy limits imposed by the device size.” Leeson goes on to analyze a simple biological coding and transmission scheme based on diffusion. This is only a start and one of the key questions remaining unanswered is the following: “Is it possible to sustain an appropriately high rate of data transmission without ‘cooking’ the brain or otherwise interfering with its normal function?”.
12 In addition to chemical synapses of the sort alluded to in the presentation, there are also electrical synapses. An electrical synapse “is a mechanical and electrically conductive link between two abutting neurons that is formed at a narrow gap between the pre- and postsynaptic neurons known as a gap junction.”
13 The presence of calcium can be used as a marker for neural activity. The method of calcium imaging is used to measure the concentrations of calcium within cells using fluorescent molecules that respond to the binding of Ca2+ ions by changing their fluorescence properties. This technique can partially reconstruct the firing patterns of relatively large populations — thousands — of neurons, but has poor temporal resolution and requires bathing the cells in light that can cause photodamage.
14 As for the even more ambitious goal of reading off the proteomic history of a behaving brain, here are some additional questions and speculations that highlight the challenges:
How do you design a retrovirus delivery vector that only targets neurons, doesn’t replicate, achieves high coverage of the target population, and can be manufactured economically in sufficient quantity?
Could you achieve this so that the encapsulated RNA instructions are identical except for a unique signature used to tag the host neuron for collecting connection attributes?
If not, could you induce the host to generate such a signature exactly once upon the first being infected, and provide some guarantee that with high probability the self-manufactured signature is unique within a given population of neurons all of which use the same method for generating their signatures?
How might we introduce the vector into the blood supply, avoid an immune response rejection, circumvent the blood-brain barrier, reliably makes its way to the host, and quietly self-destruct if the host is already infected?
Are there existing options for retroviruses that are effectively benign allowing the host to perform normally after the initial infection and ward off subsequent infections to avoid altering the signature?
How can we control the rate at which the new molecular machinery propagates information packets across synapses, or perhaps we can tag the neurotransmitters so that they convey the signature of the transmitting neuron?
What are some candidate cellular machines that could be adapted to sense neural states and how might they be modified to carry out sensing operations without interfering with their normal function?
Could you alter a ribosome so that as a side effect of transcription it also produces a marker for the particular protein — perhaps it could do this some fraction of the time such that the marked proteins are proportional to total production?
If not, could a completed protein be tagged — perhaps in an epigenetic fashion via methylation, or might it be better to tag and package the mRNA after it has performed its purpose assuming that the process leaves the single-stranded mRNA intact?
Once the data has been recorded and packaged to include the necessary provenance to enable reconstruction of the neural-state information, and is either floating free within the cell membrane or secured to some organelle as a staging area for subsequent post processing, how would you perform any additional protective cloaking and transfer the packaged information outside the cell body and subsequently into the blood-lymph system?
